-
Network Design Principles
Classify Common Molecular AlgorithmsWhat are the core regulatory algorithms (both dynamic and spatial) needed to build a living cell?
Given biochemical constraints, are their finite ways to achieve these functions using networks built from biological molecules (i.e. a “periodic table” of common biological algorithms)?
Design principles of regulatory networks: searching for the molecular algorithms of the cell (Mol. Cell, 2013)
-
Reconstituting Signaling Systems
Can we reconstitute signaling pathways and networks to uncover their mechanisms of signal transmission, specificity and information processing? We reconstitute systems both in vitro (biochemical reconstitution), as well as in vivo using heterologous engineered cellular systems (synthetic reconstitution).
Reconstitution allows us to systematically and surgically vary individual system parameters to map precisely how these parameters contribute to system function and fitness. Tools such as optogenetics give us dials and knobs with which we can precisely perturb these complex systems in living cells.
Build life to understand it
(Nature 2010)Rewiring cells: synthetic biology as a tool to interrogate the organizational principles of living systems
(Ann. Rev Biophys, 2010)Scaffold proteins: hubs for controlling the flow of cellular information
(Science 2011) -
Evolution and Design of Signaling Proteins
How do new signaling functions and connectivity arise, and what is the role of modularity in evolution?
Can we map and understand the evolution of signaling proteins and networks by comparing the sequences, composition, and function interoperability of proteins from related species?
Can we understand the functional plasticity and evolvability of signaling proteins and networks by attempting to engineer or combinatorially select for new function?
Can we design novel receptors for disease related signals?
Phosphotyrosine signaling: evolving a new cellular communication system
(Cell 2010)Exploitation of Latent Allostery Enables the Evolution of New Modes of MAP Kinase Regulation
(Cell 2013)Rapid diversification of cell signaling phenotypes by modular domain recombination
(Science 2010) -
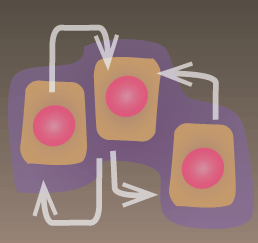
Collective Decisions & Self-organization
Most cells make important decisions through a combination of both autonomous (individual cell) responses and collective responses, mediated by secretion-based cell-cell communication (e.g. autocrine, paracrine, quorum sensing).
Currently, it is extremely challenging to distinguish autonomous and collective responses. We are using synthetic signaling systems and orthogonal cell-cell communication systems to disentangle these circuits. We hope to determine common ways in which cell-cell signaling can process and reshape autonomous responses.
We are also trying to understand how cell-cell communication network motifs contribute to emergent multicellular spatial self-organization in processes such as development.
Secreting and sensing the same molecule allows cells to achieve versatile social behaviors
(Science 2014)Designing synthetic regulatory networks capable of self-organizing cell polarization (Cell 2012)
-

Redesigning the T cell
Custom Therapeutics CellsWe are applying our knowledge of signaling networks to engineer cells with custom therapeutic responses. We are focusing on engineering T cells, as T cells armed with synthetic chimeric antigen receptors (CARs) have proven to be a highly effective clinical platform for recognizing and killing cancer cells. We are designing next generation therapeutic cells that are safer yet more potent by detecting and integrating multiple control inputs. We have received an NIH Director’s Transformative RO1 grant for this project. We are also interested in engineering therapeutic cells for autoimmunity, infectious disease, and regenerative medicine.
Cell-based therapeutics: the next pillar of medicine
(Science Trans. Med 2013)Designing customized cell signaling circuits
(Nature Rev. Mol. Cell Bio. 2010) -

Cellular Optogenetics
Interrogating Spatial & Dynamic ControlWe have developed light-responsive protein interaction modules that give us a remarkable ability to perturb and manipulate intracellular signaling nodes in a precise spatial and temporal manner. We use these tools to probe how cells encode and decode spatial and dynamic information.
We also use optogenetic profiling of cancer cells to quantitatively map functional changes in their growth signaling networks. Such analysis may uncover key mechanisms involved in cancer, and may provide novel ways to classify and treat cancer types.
The promise of optogenetics in cell biology: interrogating molecular circuits in space and time
(Nature Meth. 2011) -
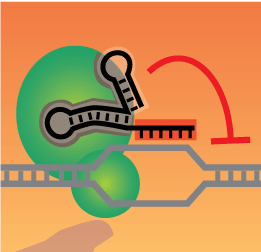
Crispr Circuits
Synthetic Regulation of the GenomeWe have developed methods to use catalytic inactive Cas9/CRISPR to regulate transcriptional control of specific endogenous genetic loci. We are using these approaches to construct novel gene regulatory circuits and cellular behaviors.
Repurposing CRISPR as an RNA-Guided Platform for Sequence-Specific Control of Gene Expression
(Cell 2013)Engineering Complex Sythetic Transcriptional Programs with CRISPR RNA Scaffolds
(Cell 2015) -

Signaling Dynamics
Using Time to Encode InformationTime is an underappreciated element in biological regulation. The dynamics of inputs (duration, frequency) can be as important as dose in controlling the magnitude and specificity of cellular responses. We are using optogenetic and microfluidic approaches to interrogate the mechanisms of dynamically regulated signaling circuits. We are also using computational and synthetic biology approaches to construct circuits that are gated by temporally encoded information.
How Do Cells Make Decisions?
Probing & Programming Cell Signaling Circuits
Living cells use genetically encoded molecular networks to monitor their environment and make sophisticated decisions. We are interested in understanding the design principles underlying the function and evolution of these networks. Cellular networks are interconnected and multifunctional; disentangling them to elucidate mechanisms is challenging. Thus, in addition to deconstructing natural networks, we use synthetic biology approaches to construct or rewire networks in ways that allow us to systematically analyze them. We study diverse cell types (yeast, T cells, neurons) to elucidate the general logic that cells use to solve common regulatory problems. As we better understand the design rules of cellular networks, we have become interested in applying this knowledge to engineer designer cells that carry out therapeutic functions, such as sensing and destroying cancer.
Our lab is composed of outstanding scientists from diverse fields (biochemistry, structural biology, cell biology, immunology, systems/synthetic biology, neurobiology, dev. biology, engineering, physics, chemistry) who are interested in applying interdisciplinary approaches to cutting edge problems in an exciting, collaborative environment.